Taxonomy
In 2021, nearly 50,000 spider species are described among 128 families. There are many times as many described insects and other invertebrates. While this may seem like a lot, numerous species remain to be described both in the wild as well as in museum collections. A scientific name, description of its appearance, short diagnosis distinguishing it from similar species, map or list of distribution records and illustrations compose a basic description, but this is virtually all biologists know about many species. The number of unknown species continues to make it an exciting frontier for naturalists, and the presence of such species in museums makes them accessible to long time specialists and undergraduate novices alike. In my academic career, I have described or redescribed 72 species among 15 genera.
Zhinu manmiaoyangi Kallal & Hormiga 2018 Phonognatha tanyodon Kallal & Hormiga 2018
Phylogeny
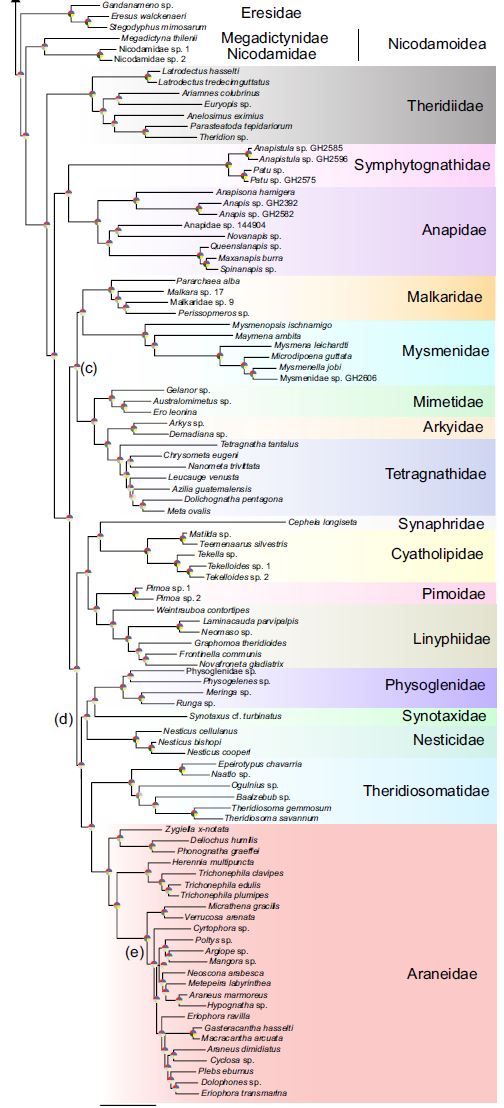
The use of increasingly sophisticated tools has been used to determine the relationships making up the spider tree of life. Ranging from coding web architectures and genitalia to analysis of thousands of protein-coding genes, the spider tree of life is still far from complete. However, many areas of the spider tree of life are coming into sharper focus thanks to the efforts of systematists around the world. I am primarily interested in the family Araneidae, which are classic orb-weaving spiders. Using morphology, behavior, and traditional loci, I was able to resolve the relationships of the early diverging araneids in the subfamily Phonognathinae. I am also part of an international team of collaborators interested in resolving the spider tree of life using next-generation sequencing which has produced a new standard for understanding spider phylogeny. The most recent of these works, published this year, includes 272 terminals. For the first time using transcriptome data, all orb-weaver families were present.
Summary topology from Kallal et al. 2020
Comparative biology
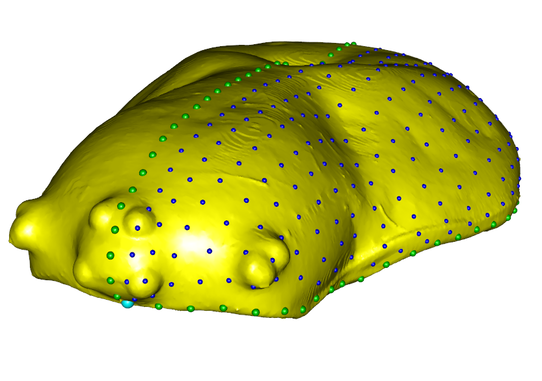
Once a supported tree is established, you can ask many questions about the biology of those organisms. These may include ancestral area estimations, which are studies in which geographic areas are coded for terminals and then areas inhabited by those terminals' ancestors are reconstructed to answer biogeographic questions. For instance, I was able to determine that member of the genera Phonognatha and Artifex colonized New Caledonia after its supposed reemergence from the sea. You could do something similar with traits, such as when I reconstructed the type of retreats used by araneoids with interest in leaf-curling phonognathines. Finally, the age of 'big data' doesn't preclude morphology from being relevant, and coding complex structures using geometric morphometrics allows you to answer questions like where orb-weaver carapace shapes are different among sexes or subfamilies, or whether they covary with size dimorphism. Most recently, I am using 3D morphometrics to analyze micro-CT reconstructions of the carapace and chelicerae of spiders to investigate modularity of the structures.
Landmarks summarizing the shape of of the carapace of Araneus marmoreus